what is the warburg method to treat cancer
Abstruse
Warburg upshot is an emerging hallmark of cancer cells with pyruvate kinase M2 (PKM2) as its key regulator. Curcumin is an extensively-studied anti-cancer compound, notwithstanding, its part in affecting cancer metabolism remains poorly understood. Herein, we evidence that curcumin inhibits glucose uptake and lactate production (Warburg effect) in a variety of cancer cell lines by downward-regulating PKM2 expression, via inhibition of mTOR-HIF1α centrality. Stable PKM2 silencing revealed that PKM2 is required for Warburg event and proliferation of cancer cells. PKM2 over-expression abrogated the furnishings of curcumin, demonstrating that inhibition of Warburg effect by curcumin is PKM2-mediated. Loftier PKM2 expression correlated strongly with poor overall survival in cancer, suggesting the requirement of PKM2 in cancer progression. The study unravels novel PKM2-mediated inhibitory result of curcumin on metabolic capacities of cancer cells. To the best of our knowledge, this is the first study linking curcumin with PKM2-driven cancer glycolysis, thus, providing new perspectives into the mechanism of its anticancer activeness.
Introduction
Metabolic priorities of cancer cells differ remarkably from normal cells, thus providing a new therapeutic window. Metabolic reprogramming in cancer cells support their growth, survival, proliferation and maintenance1. In 1920's, Otto Warburg observed that tumor cells produce big quantities of lactate even when sufficient oxygen is nowadays, a phenomenon referred to as Warburg effect or aerobic glycolysis ii. Warburg effect is characterized by high glucose uptake and lactate release and is now considered as a hallmark of nearly all tumorsthree. This metabolic adaptation benefits cancer cells in surviving through hypoxic atmospheric condition, commonly found in tumors, and to support their anabolic requirements4,v. Tendency of cancer cells to take-upwardly big quantities of glucose is exploited in clinical detection of tumors by 18fluorodeoxyglucose-positron emission tomography (FDG-PET) scan6. Hyper-activating mutations in growth signaling are known to induce expression of enzymes that are vital for cancer metabolism7. Interest in studying cancer metabolism has been rekindled recently with a outburst in number of publications in concluding decade8. Now, metabolism of cancer cells is considered a therapeutic hotspot for dietary and pharmacologic interventions9,10,xi.
All-encompassing studies have shown that pyruvate kinase M2 (PKM2) is 1 of the critical regulator of Warburg upshot12,13. Accordingly, PKM2 is highly expressed in proliferating cells like tumor and embryonic cellsfourteen. PKM2 is ane of the four isoform of pyruvate kinase- PKL, PKR, PKM1 and PKM215. Switch towards PKM2 isoform is a pre-requisite for aerobic glycolysis to take place16. Oncogenic transcription cistron c-Myc has been demonstrated to control the mutually exclusive alternative splicing of PKM mRNA in favor of PKM217. PKM2 expression has been used as a tumor marker18,19,20,21. Nosotros and others have previously reported that PKM2 is associated with tumor metabolism and growth22,23. Several studies have suggested PKM2 as a therapeutic target for cancer handling24,25. Therefore, it is pertinent to evaluate drugs that could suppress PKM2 expression, thus, inhibiting cancer metabolism.
Curcumin (diferuloylmethane) is a well-known phytopolyphenolic compound isolated from rhizome of the establish Curcuma longa (Zingiberaceae)26. Curcumin is considered every bit a valuable medicinal plant in Indian systems of medicine. Numerous studies have shown the anti-cancer properties of curcumin in a wide variety of cell lines and animals27,28,29,30,31,32,33. The major features of carcinogenesis have been shown to be inhibited by curcumin34. Several mechanisms for anti-cancer activities of curcumin take been proposed, including, consecration of apoptosis34, p53 stabilization35, mTOR33, Wnt36, Notch37, PI3K38, signaling inhibition, AMPK activation39, cell wheel inhibitionforty, inhibition of oncogenes41, inactivation of NF-kB42, metastasis inhibition43, angiogenesis inhibition44, miRNA regulation45, Dna impairment and repair46. However, the event of curcumin on cancer metabolism, an emerging hallmark of cancer, remains unknown.
Here, we investigated the effect of curcumin on cancer metabolism and study novel PKM2-mediated inhibitory effects of curcumin on Warburg effect. Our results identify a new anti-cancer mechanism of curcumin and endorse its therapeutic relevance in inhibiting cancer.
Results
Curcumin inhibits Warburg effect in cancer cells
The effect of curcumin on Warburg issue was studied by measuring the rate of glucose uptake and lactate product in cancer cell lines- lung (H1299), breast (MCF-7), cervical (HeLa) and prostate (PC3) and human embryonic kidney (HEK) 293 cells, taken as control. Sub-toxic concentrations of 0–twenty μM curcumin for 24 hours were used for the study. Meaning inhibition in glucose uptake and lactate release was observed across the 4 prison cell lines, however, no observable decrease in Warburg effect was observed in HEK 293 cells (Fig. 1a,b). Dose-dependent subtract in Warburg effect started at 2.v μM with maximal subtract at 20 μM curcumin.
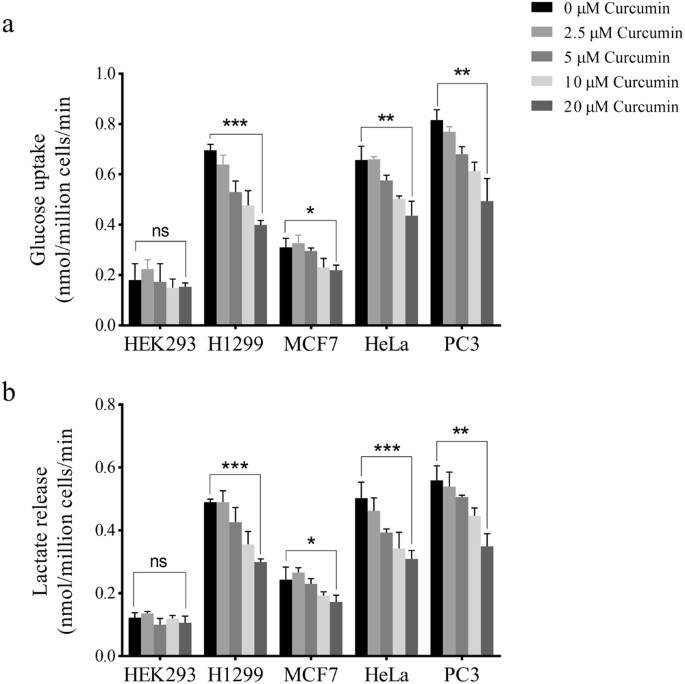
Dose-dependent outcome of curcumin on Warburg effect. High glucose uptake and lactate product, also referred to as Warburg effect, is a authentication characteristic of cancer cells (encounter text) needed to support proliferation of cancer cells. Glucose uptake (a) and lactate release (b) by H1299, MCF-seven, HeLa and PC3 cells was reduced significantly upon curcumin treatment compared with HEK293 where the change in glucose and lactate was not meaning. Different doses of curcumin (two.five, five, 10 and 20 μM) for 24 hours were used for treatment purpose. Maximal decrease in Warburg upshot was observed at 20 μM. Mistake bars stand for mean ± SD.
Curcumin down-regulates PKM2 via inhibition of mTOR-HIF1α centrality
To understand the decrease in glucose consumption and lactate product by curcumin-treated cell lines, nosotros studied the status of PKM2, a critical regulator of Warburg result. Since maximal subtract in Warburg effect was observed at 20 μM curcumin, we used this concentration to written report the upshot of curcumin on PKM2 status in H1299, MCF-7, HeLa and PC3 cell lines. Curcumin treatment substantially reduced PKM2 mRNA and poly peptide as assessed by qRT-PCR and immunoblotting (Fig. 2a,b and Supplementary Figure S2). Farther, in an attempt to elucidate the mechanism responsible for PKM2 downwardly-regulation, we studied the mTOR/HIF1α pathway inhibition upon curcumin treatment. mTOR is frequently hyper-activated in various cancers47 and curcumin has been shown to inhibit mTOR signaling48. HIF1α is a known transcriptional activator of PKM213,49. Upon curcumin treatment, decreased PKM2 expression coincided with decreased Threonine 389 (T389) phosphorylation of p70S6 kinase and decreased HIF1α protein, suggesting that curcumin down regulated PKM2 by inhibiting the mTOR/HIF1α signaling. In addition, inhibition of PKM2 expression by rapamycin (a well-known mTOR inhibitor), farther validated that curcumin decreased PKM2 via inhibition of mTOR/HIF1α signaling (Fig. 2c). GLUT1 and HKII mRNA were likewise plant to be decreased upon curcumin treatment, suggesting the contribution of these enzymes, in add-on to PKM2, in inhibition of Warburg effect upon curcumin treatment (Supplementary Figure S1).
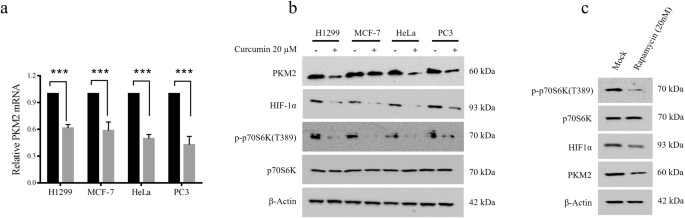
PKM2 expression is inhibited by curcumin through mTOR/HIF1α axis. (a) Substantial reduction in PKM2 mRNA in H1299, MCF-7, HeLa and PC3 cells treated with 20 μM for 24 hours. (b) Immunoblot of H1299, MCF-7, HeLa and PC3 cells showing coincided reduction in PKM2, HIF1α and phosphorylated-p70S6K (T389), upon curcumin treatment. (c) Treatment with standard mTOR inhibitor-rapamycin also showed similar expression pattern. Results suggested that curcumin inhibited PKM2 expression via inhibition of mTOR and HIF1α. Error bars in PKM2 mRNA graph represent mean ± SD.
Curcumin decreases viability of cancer cells
To analyze if inhibition of PKM2 and cancer metabolism by curcumin contributed to reduction in viability, growth of HEK293, H1299, MCF-7, HeLa and PC3 cells was assessed over the period of 24–72 hours in presence of xx μM curcumin. Decreasing trend of viability was observed in a time dependent mode (Fig. 3a). Maximum decrease in viability of all jail cell lines was observed at 72 hours, although; driblet in viability started at 24 hours. However, no significant decrease in viability was observed in control HEK 293 cells.
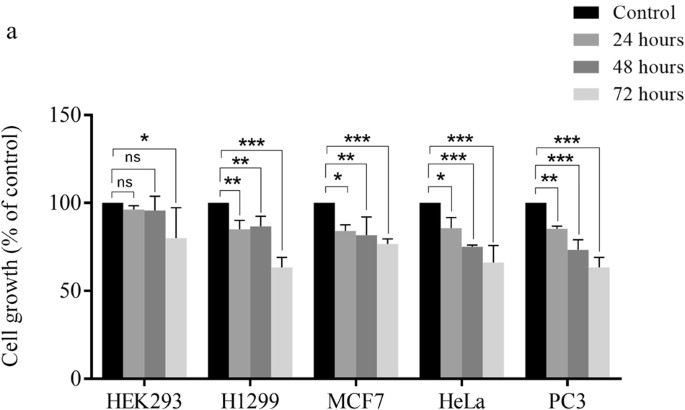
Decreased jail cell viability upon curcumin treatment. (a) Curcumin handling decreased viability of H1299, MCF-7, HeLa and PC3 cells over the fourth dimension course of 24, 48 and 72 hours. Decrease in viability of four cancer cell lines is significantly college compared to control HEK293 cells. Maximum decrease in viability of all studied cell lines was observed at 72 hours. Error bars stand for mean ± SD.
PKM2 silencing decreases Warburg effect and cell viability
Nosotros conjectured that the decrease in Warburg effect upon curcumin handling is, at least in part, due to downwardly-regulation of PKM2 expression. To this stop, PKM2 was stably silenced in H1299 cells using shRNA arroyo. Knock-down efficiency was checked by Western blotting (Fig. 4a). Thereafter, glucose consumption and lactate release were measured. shPKM2 transfected H1299 cells exhibited significant reduction in consumption of glucose and lactate product, indicating that PKM2 is crucial for Warburg effect (Fig. 4b,c). These results validated that curcumin inhibited aerobic glycolysis by down-regulating PKM2 expression. In addition to Warburg issue, PKM2 silencing also reduced viability of H1299 cells (Fig. 4d), further suggesting that PKM2-driven Warburg effect is essential for survival of cancer cells.
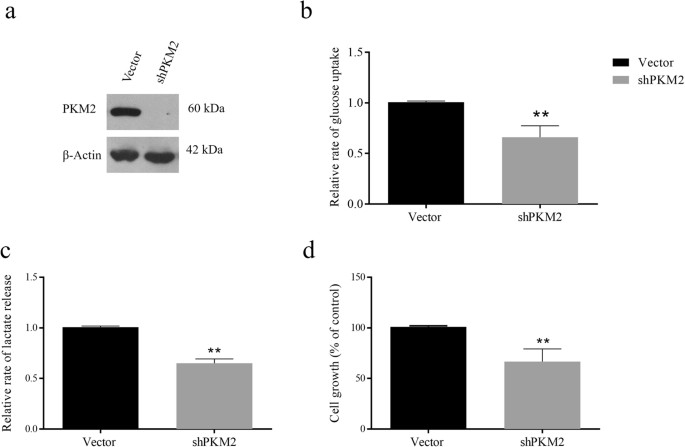
Knock-down of PKM2 reduces glucose uptake, lactate production and cell viability. PKM2 was stably silenced in H1299 cells using lentiviral shRNA as described in methods. (a) Confirmation of PKM2 knockdown by Western blotting, (b) drop in glucose uptake, (c) lactate production and (d) cell viability was observed. Error bars in PKM2 mRNA graph represent mean ± SD.
Over-expressing PKM2 reverses the effect of curcumin on metabolism of cancer cells
In society to further validate that inhibitory effects of curcumin on cancer glycolysis are PKM2-mediated, PKM2 was transiently over-expressed in H1299 cells in absence and presence of 20 μM curcumin. H1299 cells were either transfected with vector command or myc-PKM2. Confirmation of transfection was washed by immunoblotting using anti-myc antibodies (Fig. 5a). Curcumin activity was measured by checking the inhibition of p70S6 kinase T389 phosphorylation. Expectedly, PKM2 over-expression resulted in augmented Warburg issue even in continuous presence of 20 μM curcumin (Fig. 5b,c). Results demonstrated the ectopic PKM2 expression repressed the effects of curcumin on glucose uptake and lactate release, substantiating that PKM2 is a target of curcumin. In absence of curcumin, PKM2 transfected cells compared with vector showed the expected increase in glucose uptake and lactate production. Consistent with Fig. 1, treated vector command showed decreased glucose and lactate compared with untreated vector command.
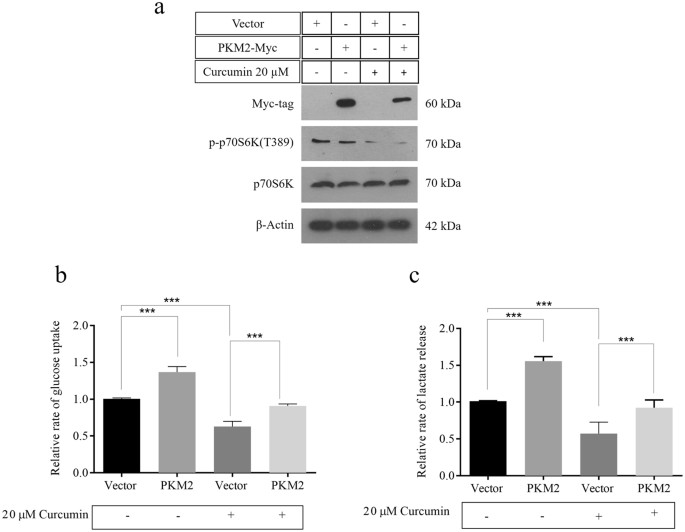
PKM2 over-expression reversed the effects of curcumin on cancer glycolysis. (a) Immunoblot confirming myc-PKM2 over-expression in H1299 cells continually exposed to 20 μM curcumin. Nearly macerated p-p70SK (T389) in both vector and PKM2 transfected H1299 cells confirmed mTOR inhibition by curcumin. Increased glucose uptake (b) and lactate release (c) in H1299 cells over-expressing PKM2, compared to vector transfected, in absence and presence of curcumin.
PKM2 expression is college in cancer and associated with poor overall survival
To investigate the expression levels of PKM2 in tumour tissues, Oncomine database was utilised. In Selamat lung cancer study comparing expression of PKM2 in normal lung (north = 58) with the lung adenocarcinoma tissue (northward = 58), PKM2 expression was found to be ii.5-fold college in lung adenocarcinoma compared to normal lung (Fig. 6a)50. Similarly, Hou dataset too revealed 2.iii-fold increase in PKM2 expression in squamous cell lung carcinoma equally compared with normal lung (n = 65) (Fig. 6a)51. Farther, we performed Kaplan-Meier analysis using online portal (world wide web.kmplot.com) to correlate PKM2 expression with overall survival of cancer patients. Higher PKM2 expression correlated strongly with poor overall survival in lung and gastric cancer patients (Fig. 6b)52,53. These results advise that higher PKM2 expression is associated with cancer and may stand for a useful prognostic marking.
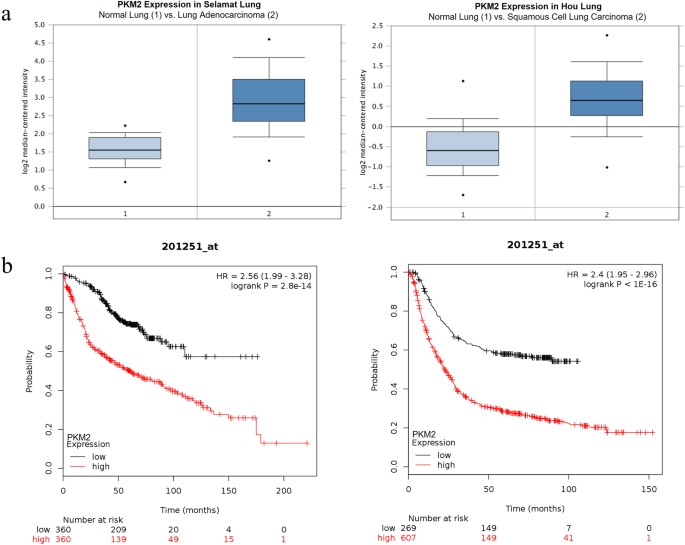
PKM2 expression is loftier in cancer and correlates with poor overall survival. (a) Box plots from Oncomine representing the college PKM2 expression in lung adenocarcinoma and squamous cell lung carcinoma, compared to normal lung. (b) Kaplan-Meier overall survival (OS) curve of patients with lung cancer expressing low and high PKM2 mRNA. (PKM2 low expression group, north = 360; PKM2 high expression grouping, northward = 360) and Kaplan-Meier overall survival (OS) curve of patients with gastric cancer expressing depression and loftier PKM2 mRNA. (PKM2 low expression group, n = 269; PKM2 high expression group, northward = 607).
Word
Metabolic transformation in cancer cells has gained enormous attending in contempo past for its immense potential equally viable therapeutic target. Owing to the metabolic vulnerabilities of cancer cells, several drugs that target cancer metabolism are under clinical trials. But every bit with any chemotherapeutic drug, drugs that target metabolism of cancer cells may have side-effects that could deteriorate patient'due south health and life-quality. On the contrary, natural plant based compounds accept relatively less toxicity and side-effects. Therefore, there is an urgent demand to screen more than natural compounds for their inhibitory effects on the emerging hallmarks of cancer, similar metabolism. Till date, very few phytochemicals have been shown to target cancer metabolism. Cleary, at that place is a need to screen more natural compounds for their negative furnishings on cancer metabolism. Metabolic enzymes could be therapeutic hotspots, provided they are key regulators of metabolic pathways operating in cancer cells. Large body of bear witness suggests that PKM2 is the key regulator of cancer metabolism, thus, targeting PKM2 to inhibit cancer metabolism should be a viable anti-cancer strategy.
Curcumin has been well-studied as a inhibitor of a diversity of cancer features; all the same, its effects on cancer metabolism remained un-elucidated. We demonstrated that curcumin inhibits cancer metabolism in a PKM2-dependent manner. Results presented here unravel a new dimension of anti-cancer mechanisms of curcumin. Results from this piece of work, along with the already known literature on curcumin, suggest that curcumin inhibits almost, if not all, hallmarks of cancer, making information technology a powerful plant-based anti-cancer chemical compound. Still, at that place are several caveats associated with the stability and bioavailability of curcumin, thus, hindering its application in clinical setting54. Owing to the unstable chemical nature of curcumin, it is known to degrade within few hours in culture media, thus, making its bioavailability abysmal55. Nonetheless, numerous studies take demonstrated the anti-cancer backdrop of curcumin56. This raises the possibility of role of the deposition products of curcumin in contributing to its onco-pharmacological effects57.
The upshot of curcumin on Warburg effect (Fig. ane) was institute to be consistent across all four cancer cell lines studied, highlighting the spectrum of curcumin as an anti-cancer drug. Also, these results as well hint at the near-universal requirement of Warburg outcome by solid cancers for their growth and survival. Interestingly, curcumin-induced inhibition of metabolism was axiomatic at a low 2.5 μM concentration, demonstrating the sensitivity of cancer cells to curcumin and dependency on aerobic glycolysis. Notably, the negligible effects of curcumin on glycolysis and viability of epithelial HEK 293 cells suggested that the observed inhibition of Warburg effect by curcumin is specific to cancer cells. The decrease in glucose consumption and lactate production impedes the growth of cancer cells because glycolytic flux provides for anabolic synthesis in cancer cells to produce macromolecules for daughter cancer cells. Furthermore, intermediates of glycolysis acts as precursors or intermediates of cross-talking anabolic pathway like pentose phosphate pathway. Also, loftier rates of glycolysis in cancer also serve the purpose of rapid ATP product, as in case of muscle during heavy exercise. Therefore, it is fathomable that high glycolytic rate is a life-line of dividing cancer cells. The decrease in PKM2 expression upon curcumin handling explains the observed decrease in aerobic glycolysis in jail cell lines treated with curcumin. Since PKM2 acts a regulator of glycolytic flux12, through its protein and pyruvate kinase activity, changes in its expression are expected to cause changes in glycolytic flux. Appropriately, silencing of PKM2 suppressed the Warburg outcome and viability (Fig. 3). Moreover, college PKM2 expression in cancer tissues and statistically significant association with poor overall survival of cancer patients, strongly suggested the part of PKM2 in cancer (Fig. 6).
mTOR is an important growth regulator and is usually very agile in cancer cells due to mutations in regulatory pathways or the mTOR itself58,59. The function of mTOR in protein synthesis makes information technology indispensable for the survival of cancer cells. The regulation of HIF1α and PKM2 expression by mTOR highlights the importance of mTOR signaling in regulation of cancer metabolism (Fig. 2). mTOR is a tuner of metabolism because latter provides raw-cloth for protein synthesis i.e. amino acids and is known to regulate HIF1α and PKM2 expression23,60,61. Therefore, inhibition of mTOR is expected to inhibit PKM2 and glycolysis, which otherwise support anabolic metabolism in cancer62. Inhibition of Warburg outcome results in decreased anabolism and therefore the viability of cancer cells is negatively affected by curcumin (Fig. 3) and silencing of PKM2 (Fig. 4) every bit all these factors converge to suppress macromolecular synthesis. The part of HIF1α in decision-making the expression of PKM2 is an outcome of hypoxic atmospheric condition that exists in tumor cells. PKM2 can drive glycolysis fifty-fifty in absence of oxygen, past promoting the conversion of pyruvate to lactate through a largely unknown mechanism, thus, induction of PKM2 by HIF1α is an accommodation necessary for tumor growth. Abrogation of inhibitory furnishings of curcumin on aerobic glycolysis in PKM2 over-expressing cells non only suggested that furnishings were mediated by PKM2 simply also highlighted the importance of PKM2 for cancer cell metabolism (Fig. 5).
Although curcumin has been shown to exert anti-cancer effects through a multifariousness of mechanisms, it is of import to identify mechanisms that could exist targeted without harming normal cells. Metabolic rewiring in cancer cells is one such mechanism. The work presented hither represents a new anti-cancer mechanism of curcumin and pin-points the enzyme responsible for mediating the anti-cancer effects of curcumin. Since bioavailability of curcumin is a claiming faced in clinical setting; stable analogues of curcumin should be tested for their ability to mimic inhibitory effects of curcumin on cancer metabolism.
In summary, this report unravels a new anti-cancer role of curcumin and invites farther research into exploiting curcumin and its analogues for successful clinical inhibition of metabolic addictions in cancer cells.
Materials and Methods
Cell culture and drug treatment
HEK293, H1299, MCF-7, HeLa and PC3 cell lines were procured and maintained as described63. Briefly, cell lines were grown in monolayer and passaged routinely 2–three times a week. Cell lines were maintained either in DMEM or RPMI with 10% FBS (Gibco) and 1% penicillin/streptomycin (Sigma) at 37°C and 5% CO2 in a humidified incubator. 2 mM Curcumin (Sigma, MO, United states) and 200 µM rapamycin (Sigma, MO, U.s.) stock solutions were prepared using DMSO and stored at −80°C, until further employ. Prior to drug treatment, cells were allowed to abound for 24 hours followed past treatment with either DMSO (mock control) or curcumin for 24–72 hours.
Metabolic assays
For glucose and lactate: Spent media was nerveless, centrifuged to remove any cell droppings and deproteinized using 5% trichloroacetic acid (TCA) and glucose and lactate were assayed using colorimetric kits (BioVision, Usa) every bit per the manufacturer's instructions. Standard curves were prepared and background corrections were washed. All measurements were normalized to jail cell numbers. Charge per unit of glucose uptake and lactate production were measured and expressed equally nmol/million cells/minute.
Western blotting and qRT-PCR
Whole cell lysate was prepared in modified RIPA buffer containing 50 mM Tris-Cl pH 7.2, 150 mM NaCl, 0.5% sodium deoxycholate, 0.1% SDS, i% Triton X-100 with added 1 mM dithiothreitol (DTT), 1 mM phenylmethylsulfonyl fluoride (PMSF), protease and phosphatse inhibitor cocktail (Geno Biosciences, USA). Lysates were and so centrifuged and supernatant containing protein was quantified using BCA (bicinchoninic acrid) kit (Thermo Scientific, Us) as per the manufacturer protocol. Equal amount of protein was loaded and separated by SDS-Folio, transferred to nitrocellulose membrane (mdi, United states of america) and probed with relevant primary antibodies. Membrane was incubated with advisable secondary antibody for i hour at room temperature and proteins were detected using Luminata Forte (Millipore, USA). Densitometry assay was carried out using ImageJ software to examine the relative cistron expression alter following normalizing with β-actin. Main antibodies used were: anti-PKM2, anti-HIF1α, anti-myc, anti-p-p70S6K (T389), anti-p70S6K and anti-β-actin (Cell signaling Engineering science, USA). For qRT-PCR: total RNA was extracted from cell lines using TRI reagent (Sigma, MO, USA) followed by reverse transcription to cDNA using Superscript® III opposite transcriptase kit (Life technologies, USA). BIORAD CFX96 TouchTM real-time machine was used for RT-PCR assay using SYBR green PCR principal mix (Practical Biosystems). Comparative C T method (2−ΔΔCT ) was used for calculation of relative factor expression. Actin was used equally endogenous command. Primers used for qRT-PCR assays are as follows: PKM2 (exon 10); forward v′-TGCAATTATTTGAGGAACTCC-3′, reverse 5′-CACTGCAGCACTTGAAGGAG-three′. HKII; forward v′- CCAACCTTAGGCTTGCCATT -3′, reverse 5′- CTTGGACATGGGATGGGGTG-3′. GLUT 1; forrard 5′- CTTTGTGGCCTTCTTTGAAGT-3′, contrary v′- CCACACAGTTGCTCCACAT-iii′. ACTIN; forward 5′- ACTCTTCCAGCCTTCCTTC-3′, reverse 5′′- ATCTCCTTCTGCATCCTGTC-three′.
Stable shRNA silencing and transient transfections
For stable knockdown of PKM2: the lentiviral particles were generated every bit described63. Briefly, HEK293T cells were transfected with transfer vector (LKO.1) containing shRNAs, plus packaging vectors- psPAX and pMD2.G, using Lipofectamine® 3000 (ThermoFisher Scientific, USA). Xl eight hours post-transfection, viral particles were harvested and used to infect the H1299 cells. Option of infected cells was washed in DMEM containing 2 µg/ml puromycin over the menstruum of two weeks. For PKM2 over-expression: pcDNA-PKM2-myc or pcDNA-myc were transfected using Lipofectamine® 3000 reagent (ThermoFisher Scientific, USA), equally per the manufacturer's instructions.
MTT analysis for cell viability
Cell viability was assessed using MTT (3-(four, v-Dimethylthiazol-2yl)-2, 5-diphenyl tetrozolium bromide) (Sigma). Exponentially growing cells were seeded in 96 well plate with the density of 10,000 cells per well. Subsequent to the cell adherence to the surface of civilisation plate (afterward approximately 12 hours), cells were treated with xx μM curcumin. Cell viability was measured every 24-hr interval (24 to 72 hours) by washing the cells and replacing with fresh medium containing 20 μl of MTT (5 mg/ml in PBS) to each well. The plates were incubated for three hours in dark. The formazon crystals developed were solubilized with 100 μl of DMSO and the plate was kept in dark for another 5–10 min. Absorbance was measured using a microplate reader (Molecular Device) at 570 nm. Cells were seeded in triplicates for each group and the experiment was independently repeated thrice.
Statistical assay
Data were represented as hateful with SD. Educatee's t-test or 1-mode or two-fashion ANOVA with multiple comparisons was used to summate statistical significance. P < 0.05 was considered to exist statistically pregnant. Statistical significance is represented every bit: *P < 0.05; **P < 0.01; ***P < 0.001.
References
-
DeBerardinis, R. J. & Chandel, Northward. South. Fundamentals of cancer metabolism. Scientific discipline advances 2, e1600200, https://doi.org/10.1126/sciadv.1600200 (2016).
-
Warburg, O., Posener, Grand. & Negelein, E. Über den Stoffwechsel der Carcinomzelle. Biochem. Zeitschr 152, 309–344, https://doi.org/10.1016/j.tibs.2015.12.001 (1924).
-
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Prison cell 144, 646–674, https://doi.org/10.1016/j.cell.2011.02.013 (2011).
-
Gatenby, R. A. & Gillies, R. J. Why practise cancers accept high aerobic glycolysis? Nature reviews. Cancer iv, 891–899, https://doi.org/10.1038/nrc1478 (2004).
-
Iqbal, M. A., Gupta, Five., Gopinath, P., Mazurek, Southward. & Bamezai, R. N. Pyruvate kinase M2 and cancer: an updated assessment. FEBS letters 588, 2685–2692, https://doi.org/ten.1016/j.febslet.2014.04.011 (2014).
-
Vander Heiden, Thousand. Thou., Cantley, L. C. & Thompson, C. B. Understanding the Warburg issue: the metabolic requirements of prison cell proliferation. Science 324, 1029–1033, https://doi.org/10.1126/scientific discipline.1160809 (2009).
-
Nagarajan, A., Malvi, P. & Wajapeyee, Northward. Oncogene-directed alterations in cancer prison cell metabolism. Trends in cancer 2, 365–377, https://doi.org/ten.1016/j.trecan.2016.06.002 (2016).
-
Liberti, Thousand. Five. & Locasale, J. West. The Warburg Effect: How Does it Benefit Cancer Cells? Trends in biochemical sciences 41, 211–218, https://doi.org/10.1016/j.tibs.2015.12.001 (2016).
-
Tennant, D. A., Duran, R. V. & Gottlieb, Due east. Targeting metabolic transformation for cancer therapy. Nature reviews. Cancer 10, 267–277, https://doi.org/x.1038/nrc2817 (2010).
-
Borriello, A. & Della Ragione, F. The new anticancer era: Tumor metabolism targeting. Prison cell Cycle 16, 310–311, https://doi.org/10.1080/15384101.2016.1271635 (2017).
-
Vernieri, C. et al. Targeting Cancer Metabolism: Dietary and Pharmacologic Interventions. Cancer discovery six, 1315–1333, https://doi.org/10.1158/2159-8290.CD-sixteen-0615 (2016).
-
Chaneton, B. & Gottlieb, Due east. Rocking cell metabolism: revised functions of the key glycolytic regulator PKM2 in cancer. Trends in biochemical sciences 37, 309–316, https://doi.org/10.1016/j.tibs.2012.04.003 (2012).
-
Prakasam, G., Iqbal, K. A., Bamezai, R. North. 1000. & Mazurek, Due south. Posttranslational Modifications of Pyruvate Kinase M2: Tweaks that BenefitCancer. Frontiers in Oncology viii, https://doi.org/10.3389/fonc.2018.00022 (2018).
-
Gupta, V. & Bamezai, R. N. Man pyruvate kinase M2: a multifunctional protein. Protein science: a publication of the Protein Guild xix, 2031–2044, https://doi.org/ten.1002/pro.505 (2010).
-
Prakasam, Chiliad., Iqbal, M. A., Gupta, V., Kumar, B. & Bamezai, R. N. K. In Encyclopedia of Snaling Molecules (ed Sangdun Choi) 1–11 (Springer New York, 2017).
-
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is of import for cancer metabolism and tumour growth. Nature 452, 230–233, https://doi.org/x.1038/nature06734 (2008).
-
David, C. J., Chen, M., Assanah, One thousand., Canoll, P. & Manley, J. Fifty. HnRNP proteins controlled past c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368, https://doi.org/10.1038/nature08697 (2010).
-
Ahmed, A. S. et al. M2-PK as a novel marker in ovarian cancer. A prospective cohort study. European journal of gynaecological oncology 28, 83–88 (2007).
-
Ewald, N. et al. Pyruvate kinase M2 (tumor M2-PK) as a screening tool for colorectal cancer (CRC). A review of current published data. Zeitschrift fur Gastroenterologie 43, 1313–1317, https://doi.org/x.1055/south-2005-858657 (2005).
-
Hardt, P. D. & Ewald, N. Tumor M2 pyruvate kinase: a tumor marker and its clinical application in gastrointestinal malignancy. Proficient review of molecular diagnostics 8, 579–585, https://doi.org/10.1586/14737159.eight.5.579 (2008).
-
Hardt, P. D. et al. Faecal neoplasm M2 pyruvate kinase: a new, sensitive screening tool for colorectal cancer. British journal of cancer 91, 980–984, https://doi.org/10.1038/sj.bjc.6602033 (2004).
-
Iqbal, G. A. et al. Missense mutations in pyruvate kinase M2 promote cancer metabolism, oxidative endurance, anchorage independence, and tumor growth in a dominant negative manner. The Journal of biological chemical science 289, 8098–8105, https://doi.org/x.1074/jbc.M113.515742 (2014).
-
Lord's day, Q. et al. Mammalian target of rapamycin up-regulation of pyruvate kinase isoenzyme blazon M2 is disquisitional for aerobic glycolysis and tumor growth. Proceedings of the National Academy of Sciences of the United States of America 108, 4129–4134, https://doi.org/10.1073/pnas.1014769108 (2011).
-
Chen, J. et al. Shikonin and its analogs inhibit cancer prison cell glycolysis past targeting tumor pyruvate kinase-M2. Oncogene thirty, 4297–4306, https://doi.org/10.1038/onc.2011.137 (2011).
-
Shi, H. S. et al. Silencing of pkm2 increases the efficacy of docetaxel in human lung cancer xenografts in mice. Cancer science 101, 1447–1453, https://doi.org/ten.1111/j.1349-7006.2010.01562.ten (2010).
-
Noorafshan, A. & Ashkani-Esfahani, S. A review of therapeutic effects of curcumin. Electric current pharmaceutical design xix, 2032–2046 (2013).
-
Ramasamy, T. S., Ayob, A. Z., Myint, H. H., Thiagarajah, South. & Amini, F. Targeting colorectal cancer stem cells using curcumin and curcumin analogues: insights into the machinery of the therapeutic efficacy. Cancer cell international 15, 96, https://doi.org/10.1186/s12935-015-0241-10 (2015).
-
Amin, A. R. et al. Curcumin induces apoptosis of upper aerodigestive tract cancer cells by targeting multiple pathways. PloS one 10, e0124218, https://doi.org/10.1371/journal.pone.0124218 (2015).
-
Radhakrishna Pillai, G., Srivastava, A. S., Hassanein, T. I., Chauhan, D. P. & Carrier, E. Induction of apoptosis in human lung cancer cells by curcumin. Cancer letters 208, 163–170, https://doi.org/10.1016/j.canlet.2004.01.008 (2004).
-
Hartojo, Due west. et al. Curcumin promotes apoptosis, increases chemosensitivity, and inhibits nuclear factor kappaB in esophageal adenocarcinoma. Translational oncology 3, 99–108 (2010).
-
Zhang, H. et al. Curcumin Promotes Cell Cycle Arrest and Inhibits Survival of Human Renal Cancer Cells by Negative Modulation of the PI3K/AKT Signaling Pathway. Prison cell biochemistry and biophysics 73, 681–686, https://doi.org/10.1007/s12013-015-0694-5 (2015).
-
Li, F., Chen, X., Xu, B. & Zhou, H. Curcumin induces p53-contained necrosis in H1299 cells via a mitochondria-associated pathway. Molecular medicine reports 12, 7806–7814, https://doi.org/10.3892/mmr.2015.4395 (2015).
-
Jiao, D. et al. Curcumin inhibited HGF-induced EMT and angiogenesis through regulating c-Met dependent PI3K/Akt/mTOR signaling pathways in lung cancer. Molecular therapy oncolytics 3, 16018, https://doi.org/ten.1038/mto.2016.18 (2016).
-
Beevers, C. Due south. et al. Curcumin disrupts the Mammalian target of rapamycin-raptor complex. Cancer research 69, 1000–1008, https://doi.org/ten.1158/0008-5472.Can-08-2367 (2009).
-
Li, Westward. et al. A preliminary study of the effect of curcumin on the expression of p53 protein in a homo multiple myeloma cell line. Oncology messages 9, 1719–1724, https://doi.org/10.3892/ol.2015.2946 (2015).
-
He, Thou. et al. Curcumin suppresses cell proliferation through inhibition of the Wnt/beta-catenin signaling pathway in medulloblastoma. Oncology reports 32, 173–180, https://doi.org/10.3892/or.2014.3206 (2014).
-
Yang, J. et al. Curcumin inhibits the survival and metastasis of prostate cancer cells via the Notch-ane signaling pathway. APMIS: acta pathologica, microbiologica, et immunologica Scandinavica 125, 134–140, https://doi.org/10.1111/apm.12650 (2017).
-
Tian, B. et al. Curcumin inhibits urothelial tumor development past suppressing IGF2 and IGF2-mediated PI3K/AKT/mTOR signaling pathway. Periodical of drug targeting, 1–eleven, https://doi.org/10.1080/1061186X.2017.1306535 (2017).
-
Kang, O. H. et al. Curcumin decreases oleic acrid-induced lipid accumulation via AMPK phosphorylation in hepatocarcinoma cells. European review for medical and pharmacological sciences 17, 2578–2586 (2013).
-
Hu, A. et al. Curcumin induces G2/M cell cycle abort and apoptosis of head and neck squamous cell carcinoma in vitro and in vivo through ATM/Chk2/p53-dependent pathway. Oncotarget, https://doi.org/x.18632/oncotarget.17096 (2017).
-
Lin, J. K. Suppression of protein kinase C and nuclear oncogene expression every bit possible activity mechanisms of cancer chemoprevention by Curcumin. Archives of pharmacal enquiry 27, 683–692 (2004).
-
Singh, Due south. & Aggarwal, B. B. Activation of transcription cistron NF-kappa B is suppressed by curcumin (diferuloylmethane) [corrected]. The Journal of biological chemistry 270, 24995–25000 (1995).
-
Liao, H., Wang, Z., Deng, Z., Ren, H. & Li, Ten. Curcumin inhibits lung cancer invasion and metastasis by attenuating GLUT1/MT1-MMP/MMP2 pathway. International periodical of clinical and experimental medicine 8, 8948–8957 (2015).
-
Bhandarkar, Southward. S. & Arbiser, J. 50. Curcumin equally an inhibitor of angiogenesis. Advances in experimental medicine and biological science 595, 185–195, https://doi.org/10.1007/978-0-387-46401-5_7 (2007).
-
Chen, J., Xu, T. & Chen, C. The disquisitional roles of miR-21 in anti-cancer effects of curcumin. Annals of translational medicine 3, 330, https://doi.org/10.3978/j.issn.2305-5839.2015.09.20 (2015).
-
Ji, Z. Targeting DNA impairment and repair by curcumin. Breast cancer: basic and clinical inquiry 4, 1–3 (2010).
-
Chen, Y., Wei, H., Liu, F. & Guan, J. L. Hyperactivation of mammalian target of rapamycin circuitous 1 (mTORC1) promotes breast cancer progression through enhancing glucose starvation-induced autophagy and Akt signaling. The Journal of biological chemistry 289, 1164–1173, https://doi.org/10.1074/jbc.M113.526335 (2014).
-
Huang, Southward. Inhibition of PI3K/Akt/mTOR signaling past natural products. Anti-cancer agents in medicinal chemistry 13, 967–970 (2013).
-
Iqbal, M. A. et al. Insulin enhances metabolic capacities of cancer cells by dual regulation of glycolytic enzyme pyruvate kinase M2. Molecular cancer 12, 72, https://doi.org/10.1186/1476-4598-12-72 (2013).
-
Selamat, South. A. et al. Genome-scale analysis of Deoxyribonucleic acid methylation in lung adenocarcinoma and integration with mRNA expression. Genome research 22, 1197–1211, https://doi.org/10.1101/gr.132662.111 (2012).
-
Hou, J. et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PloS one 5, e10312, https://doi.org/x.1371/periodical.pone.0010312 (2010).
-
Gyorffy, B., Surowiak, P., Budczies, J. & Lanczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-jail cell lung cancer. PloS i 8, e82241, https://doi.org/10.1371/journal.pone.0082241 (2013).
-
Szasz, A. M. et al. Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic information of 1,065 patients. Oncotarget 7, 49322–49333, https://doi.org/10.18632/oncotarget.10337 (2016).
-
Shen, L., Liu, C. C., An, C. Y. & Ji, H. F. How does curcumin work with poor bioavailability? Clues from experimental and theoretical studies. Scientific reports 6, 20872, https://doi.org/ten.1038/srep20872 (2016).
-
Nelson, K. Thousand. et al. The Essential Medicinal Chemistry of Curcumin. Journal of medicinal chemistry 60, 1620–1637, https://doi.org/ten.1021/acs.jmedchem.6b00975 (2017).
-
Heger, 1000., van Golen, R. F., Broekgaarden, M. & Michel, K. C. The molecular basis for the pharmacokinetics and pharmacodynamics of curcumin and its metabolites in relation to cancer. Pharmacological reviews 66, 222–307, https://doi.org/10.1124/pr.110.004044 (2014).
-
Shen, L. & Ji, H. F. The pharmacology of curcumin: is it the degradation products? Trends in molecular medicine eighteen, 138–144, https://doi.org/10.1016/j.molmed.2012.01.004 (2012).
-
Grabiner, B. C. et al. A various assortment of cancer-associated MTOR mutations are hyperactivating and tin predict rapamycin sensitivity. Cancer discovery iv, 554–563, https://doi.org/10.1158/2159-8290.CD-thirteen-0929 (2014).
-
Yuan, T. L. & Cantley, L. C. PI3K pathway alterations in cancer: variations on a theme. Oncogene 27, 5497–5510, https://doi.org/10.1038/onc.2008.245 (2008).
-
State, S. C. & Tee, A. R. Hypoxia-inducible factor 1alpha is regulated past the mammalian target of rapamycin (mTOR) via an mTOR signaling motif. The Periodical of biological chemical science 282, 20534–20543, https://doi.org/10.1074/jbc.M611782200 (2007).
-
Hudson, C. C. et al. Regulation of hypoxia-inducible cistron 1alpha expression and function past the mammalian target of rapamycin. Molecular and cellular biology 22, 7004–7014 (2002).
-
Laplante, 1000. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293, https://doi.org/10.1016/j.prison cell.2012.03.017 (2012).
-
Prakasam, G. et al. Pyruvate kinase M knockdown-induced signaling via AMP-activated protein kinase promotes mitochondrial biogenesis, autophagy, and cancer cell survival. The Journal of biological chemical science 292, 15561–15576, https://doi.org/10.1074/jbc.M117.791343 (2017).
Acknowledgements
Department of Science and Applied science (DST), Govt. of India, for DST-INSPIRE faculty accolade and enquiry grant (DST/INSPIRE/04/2015/000556) to MAI.
Writer information
Affiliations
Contributions
Information acquisition: F.A.S., P.G., S.C., A.U.R., R.A.P., Thou.A.A., R.I., M.M.; Data assay and interpretation: M.A.I., F.A.S., P.Yard.; Wrote the manuscript: M.A.I.; Contributed reagents/analytical tools: K.A.I., R.Due north.G.B., Chiliad.H., S.M.A.; Conception, design and supervision of study: Yard.A.I. All authors contributed to manuscript revision.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary fabric
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits apply, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original writer(s) and the source, provide a link to the Creative Commons license, and indicate if changes were fabricated. The images or other tertiary party material in this commodity are included in the commodity'south Creative Eatables license, unless indicated otherwise in a credit line to the cloth. If textile is not included in the article'due south Creative Commons license and your intended use is not permitted past statutory regulation or exceeds the permitted use, y'all will need to obtain permission straight from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Reprints and Permissions
About this article
Cite this article
Siddiqui, F.A., Prakasam, G., Chattopadhyay, S. et al. Curcumin decreases Warburg outcome in cancer cells past downwardly-regulating pyruvate kinase M2 via mTOR-HIF1α inhibition. Sci Rep 8, 8323 (2018). https://doi.org/10.1038/s41598-018-25524-3
-
Received:
-
Accepted:
-
Published:
-
DOI : https://doi.org/10.1038/s41598-018-25524-iii
Further reading
-
Metabolic changes in triple negative breast cancer-focus on aerobic glycolysis
Molecular Biology Reports (2021)
-
Elevated expression of RUNX3 co-expressing with EZH2 in esophageal cancer patients from Bharat
Cancer Cell International (2020)
-
Herbal nutraceuticals: condom and potent therapeutics to battle tumor hypoxia
Periodical of Cancer Research and Clinical Oncology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you detect something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.
Source: https://www.nature.com/articles/s41598-018-25524-3
0 Response to "what is the warburg method to treat cancer"
Enregistrer un commentaire